Note
Go to the end to download the full example code.
Fine-tuning a pre-trained model¶
Warning
Finetuning is currently only available for the PET architecture.
This is a simple example for fine-tuning PET-MAD (or a general PET model), that
can be used as a template for general fine-tuning with metatrain.
Fine-tuning a pretrained model allows you to obtain a model better suited for
your specific system. You need to provide a dataset of structures that have
been evaluated at a higher reference level of theory, usually DFT. Fine-tuning
a universal model such as PET-MAD allows for reasonable model performance even if little
training data is available.
It requires using a pre-trained model checkpoint with the mtt train command and
setting the new targets corresponding to the new level of theory in the options.yaml
file.
We start by setting up the options.yaml file. Here we specify to fine-tune on a
small model dataset containing structures of ethanol, labelled with energies and
forces. We can specify the fine-tuning method in the finetune block in the
training options of the architecture. Here, the basic full option is
chosen, which finetunes all weights of the model. All available fine-tuning methods
are found in the concepts page Fine-tuning. This
section discusses implementation details, options and recommended use cases. Other
fine-tuning options can be simply substituted in this script, by changing the
finetune block.
Note
Since our dataset has energies and forces obtained from reference calculations,
different from the reference of the pretrained model, it is recommended to create a
new energy head. Using this so-called energy variant can be simply invoked by
requesting a new target in the options file. Follow the nomenclature
energy/{yourname}.
Furthermore, you need to specify the checkpoint, that you want to fine-tune in
the read_from option.
A simple options-ft.yaml file for this task could look like this:
architecture:
name: pet
training:
batch_size: 8
num_epochs: 10
learning_rate: 1e-3
warmup_fraction: 0.01
finetune:
method: full
read_from: pet-mad-v1.1.0.ckpt
training_set:
systems:
read_from: ethanol_reduced_100.xyz
reader: ase
length_unit: angstrom
targets:
energy/finetune:
quantity: energy
read_from: ethanol_reduced_100.xyz
reader: ase
key: energy
unit: eV
description: pbe energy ethanol
forces:
read_from: ethanol_reduced_100.xyz
reader: ase
key: forces
validation_set: 0.1
test_set: 0.1
In this example, we specified a low number of num_epochs and a relatively high
learning_rate, for short compilation time. Usually, the learning_rate is
chosen to be relatively low. Typically lower, than the learning_rate that the model
has been per-trained on.
to stabilise training.
Warning
Note that in targets we use the energy/finetune head, differing from the
default energy head. This means, that the model creates a new head with a new
composition model for the new reference energies provided in your dataset. While
the old energy reference is still available, it is rendered useless, as we trained
all weights of the model. If you want to obtain a model with multiple energy heads,
you can simply train on multiple energy references simultaneously. This and other
more advanced fine-tuning strategies are discussed in
Fine-tuning concepts.
We assumed that the pre-trained model is trained on the dataset
ethanol_reduced_100.xyz in which energies are written in the energy key of
the info dictionary of the dataset.
Additionally, forces should be provided with corresponding keys
which you can specify in the options-ft.yaml file under targets.
Further information on specifying targets can be found in the data section of
the Training YAML Reference.
Note
It is important that the length_unit is set to angstrom and the energy
unit is eV in order to match the units of your reference data.
After setting up your options-ft.yaml file, you can then simply run:
mtt train options-ft.yaml -o model-ft.pt
You can check finetuning training curves by parsing the train.csv that is written
by mtt train. We remove the old outputs folder from other examples, which
is not necessary for the normal usage.
import glob
import subprocess
import ase.io
import matplotlib.pyplot as plt
import numpy as np
from metatomic.torch.ase_calculator import MetatomicCalculator
# In order to obtain a pretrained model, you can use a PET-MAD checkpoint from
# huggingface. Here, we get the PET-MAD ckpt, run ``mtt train`` as a subprocess, and
# delete the old outputs folder and old model checkpoints.
subprocess.run(["rm", "-rf", "outputs"], check=True)
subprocess.run(
[
"wget",
"https://huggingface.co/lab-cosmo/pet-mad/resolve/v1.1.0/models/pet-mad-v1.1.0.ckpt",
],
check=True,
)
subprocess.run(["mtt", "train", "options-ft.yaml", "-o", "model-ft.pt"], check=True)
subprocess.run(["rm", "-rf", "pet-mad-v1.1.0.ckpt"], check=True)
CompletedProcess(args=['rm', '-rf', 'pet-mad-v1.1.0.ckpt'], returncode=0)
After training, we can check if finetuning was successful.
First we check the training curves, that are saved in train.csv in the outputs
folder. We start with parsing the csv file.
csv_path = glob.glob("outputs/*/*/train.csv")[-1]
with open(csv_path, "r") as f:
header = f.readline().strip().split(",")
f.readline() # skip units row
# Build dtype
dtype = [(h, float) for h in header]
# Load data as plain float array
data = np.loadtxt(csv_path, delimiter=",", skiprows=2)
# Convert to structured
structured = np.zeros(data.shape[0], dtype=dtype)
for i, h in enumerate(header):
structured[h] = data[:, i]
Now, let’s plot the learning curves.
training_energy_RMSE = structured["training energy/finetune RMSE (per atom)"]
training_forces_MAE = structured["training forces[energy/finetune] MAE"]
validation_energy_RMSE = structured["validation energy/finetune RMSE (per atom)"]
validation_forces_MAE = structured["validation forces[energy/finetune] MAE"]
fig, axs = plt.subplots(1, 2, figsize=((8, 3.5)))
axs[0].plot(training_energy_RMSE, label="training energy RMSE")
axs[0].plot(validation_energy_RMSE, label="validation energy RMSE")
axs[0].set_xlabel("Epochs")
axs[0].set_ylabel("energy / meV")
axs[0].set_xscale("log")
axs[0].set_yscale("log")
axs[0].legend()
axs[1].plot(training_forces_MAE, label="training forces MAE")
axs[1].plot(validation_forces_MAE, label="validation forces MAE")
axs[1].set_ylabel("force / meV/A")
axs[1].set_xlabel("Epochs")
axs[1].set_xscale("log")
axs[1].set_yscale("log")
axs[1].legend()
plt.tight_layout()
plt.show()
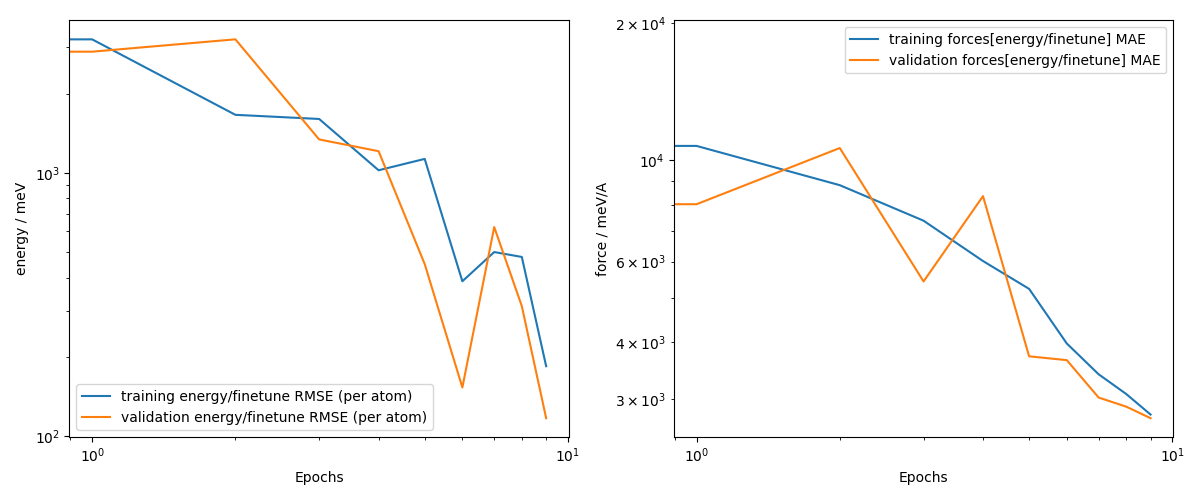
You can see that the validation loss still decreases, however, for the sake of brevity
of this exercise we only finetuned for a few epochs. As further check for how well
your fine-tuned model performs on a dataset of choice, we can check the parity plots
for energy and force
(see Model validation with parity plots).
For evaluation, we can compare performance of our fine-tuned model and the base model
PET-MAD. Using mtt eval we can simply evaluate our new energy head, by specifying
it in the options-ft-eval.yaml:
systems: ethanol_reduced_100.xyz
targets:
energy/finetune:
key: energy
unit: eV
forces:
key: forces
and then run
mtt eval model-ft.pt options-ft-eval.yaml -o output-ft.xyz
Then you can simply read the predicted energies in the headers of the xyz file.
Another possibility is to load your fine-tuned model model-ft.pt as metatomic
model and evaluate energies and forces with ASE in Python.
sphinx_gallery_capture_repr_block = ()
np.seterr()
targets = ase.io.read(
"ethanol_reduced_100.xyz",
format="extxyz",
index=":",
)
calc_ft = MetatomicCalculator(
"model-ft.pt", variants={"energy": "finetune"}, extensions_directory=None
)
# specify variant suffix here
with np.errstate(invalid="ignore"):
e_targets = np.array(
[frame.get_total_energy() / len(frame) for frame in targets]
) # target energies
f_targets = np.array(
[frame.get_forces().flatten() for frame in targets]
).flatten() # target forces
for frame in targets:
frame.calc = calc_ft
e_predictions = np.array(
[frame.get_total_energy() / len(frame) for frame in targets]
) # predicted energies
f_predictions = np.array(
[frame.get_forces().flatten() for frame in targets]
).flatten() # predicted forces
fig, axs = plt.subplots(1, 2, figsize=(8, 4))
# Parity plot for energies
axs[0].scatter(e_targets, e_predictions, label="FT")
axs[0].axline((np.min(e_targets), np.min(e_targets)), slope=1, ls="--", color="red")
axs[0].set_xlabel("Target energy / meV")
axs[0].set_ylabel("Predicted energy / meV")
min_e = np.min(np.array([e_targets, e_predictions])) - 2
max_e = np.max(np.array([e_targets, e_predictions])) + 2
axs[0].set_title("Energy Parity Plot")
axs[0].set_xlim(min_e, max_e)
axs[0].set_ylim(min_e, max_e)
# Parity plot for forces
axs[1].scatter(f_targets, f_predictions, alpha=0.5, label="FT")
axs[1].axline((np.min(f_targets), np.min(f_targets)), slope=1, ls="--", color="red")
axs[1].set_xlabel("Target force / meV/Å")
axs[1].set_ylabel("Predicted force / meV/Å")
min_f = np.min(np.array([f_targets, f_predictions])) - 2
max_f = np.max(np.array([f_targets, f_predictions])) + 2
axs[1].set_title("Force Parity Plot")
axs[1].set_xlim(min_f, max_f)
axs[1].set_ylim(min_f, max_f)
fig.tight_layout()
plt.show()
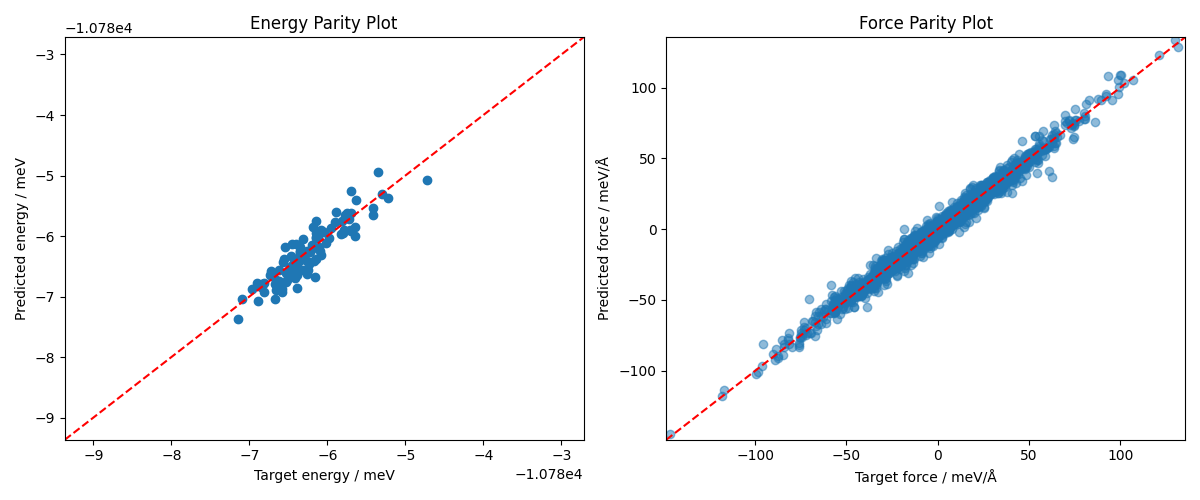
We see that the fine-tuning gives reasonable predictions on energies and forces. Since the training was limited to 10 epochs in this example, the results can be obviously improved by training for more epochs and optimizing training hyperparameters.
Note
To learn about more elaborate fine-tuning strategies and tools, check out the fine-tuning examples in the AtomisticCookbook
Total running time of the script: (0 minutes 34.443 seconds)